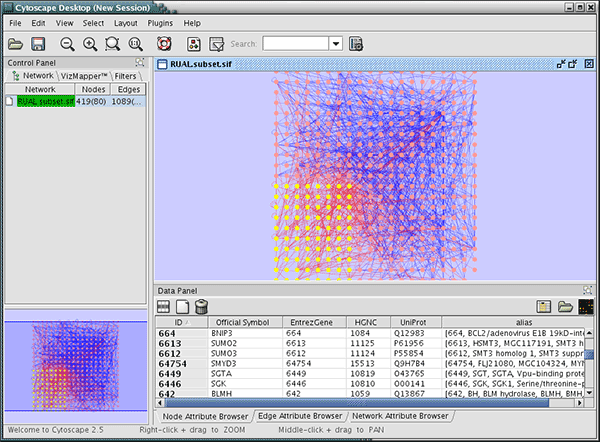
Options to rename, remove, create and copy a Style, and an option to This will reveal a set of buttons for selecting and deleting styles. The list of available styles can be edited by clicking the Edit. The Show only Applied Styles button will filter the list to display only the applied style. The list of available styles is searchable using the search field. With a specific style selected, the Style panel displays theĭetails for a given style and is used to edit these details as well.Īt the top of the interface, there is a drop-down for selecting Styles using the drop-down and the Options menu. 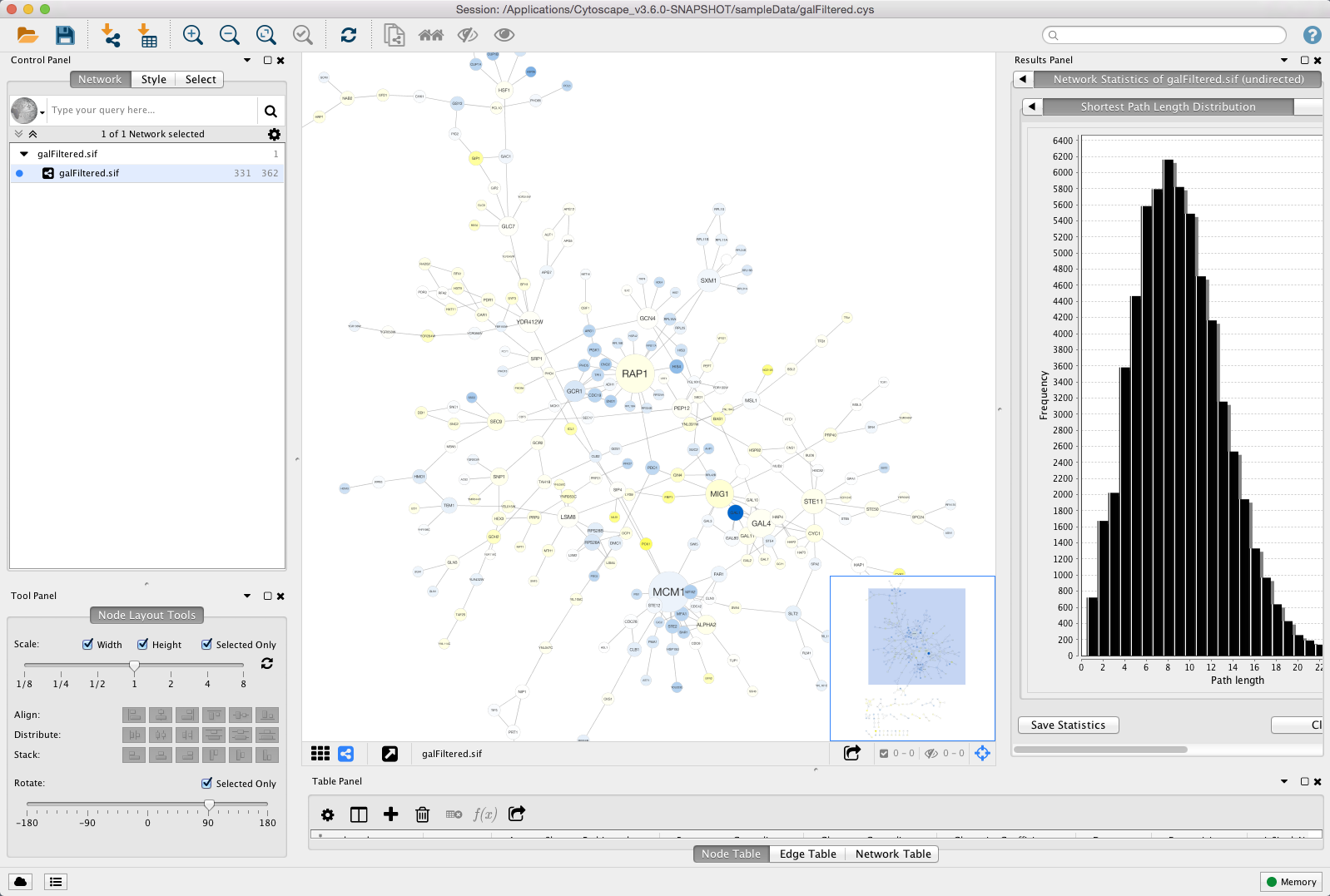 
This interface allows you to create/delete/view and switch between different The Style interface is located under the Style panel of the Show highly-connected region by edge bundling and opacity.Īdd photo/image/graphics on top of nodes.Ĭytoscape 3 has several sample styles. Use specific line types to indicate different types of interactions.Ĭontrol edge transparency (opacity) using edge weights.Ĭontrol multiple edge properties using edge score.īrowse extremely-dense networks by controlling the opacity of nodes. Visualize gene expression data along a color gradient.Įncode specific physical entities as different node shapes. or, set the font size of the node labels instead. Set node sizes based on the degree of connectivity of the nodes. Specify a default color and shape for all nodes. Interface, the appearance of your network is easily customized. Mapped table data sets is called a Style and can be created orĮdited in the Style panel of the Control Panel. Transparency, or font type) of the network. Under Available for Install » Network Inference, select GeneMANIA.One of Cytoscape's strengths in network visualization is the ability toĪllow users to encode any table data (name, type, degree, weight,Įxpression data, etc.) as a property (such as color, size of node,.In Cytoscape, go to apps » Manage apps.Adjust the amount of memory allocated to Cytoscape to at least 1.5GB.Note: Queries that use more than 1.5 GB of RAM require Cytoscape to be launched with a 64-bit JVM Java 1.5+ JVM (1.6 is required for Cytoscape 2.8+).Montojo J, Zuberi K, Rodriguez H, Kazi F, Wright G, Donaldson SL, Morris Q, Bader GD (2010)įull Article ( PDF) Minimum System Requirements 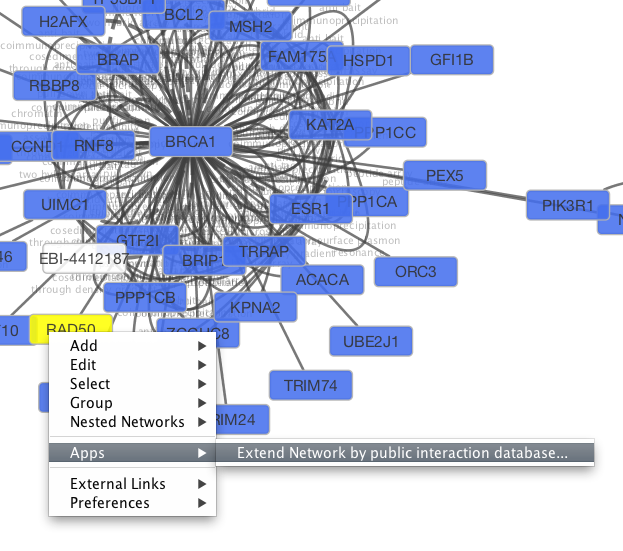
GeneMANIA Cytoscape app: fast gene function predictions on the desktop. Also, results can be saved in Cytoscape session files for later work, backup or sharing with colleagues. Integration with the popular Cytoscape network visualization and analysis platform so Cytoscape networks can be imported into GeneMANIA and GeneMANIA results can be used in other Cytoscape analysis.Powerful command line tools to automate basic and advanced analysis not available via the website.Number of query genes is limited only by the amount of memory available.The app follows the look and feel of the GeneMANIA website, but provides more features for power-users: Users may add their own interaction networks and expression profile data to complement or override the default data. The app uses a large database of functional interaction networks from multiple organisms and each related gene is traceable to the source network used to make the prediction. GeneMANIA identifies the most related genes to a query gene set using a guilt-by-association approach. The GeneMANIA Cytoscape app brings fast gene function prediction capabilities to the desktop.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
